Nucleic Acid Electrophoresis Reagents
Nucleic acid electrophoresis is a method for separating DNA and RNA, which uses the negatively charged property of nucleic acid to move it through agarose gel and the like toward an anode. It is used for confirming the yield and analyzing the size of nucleic acid obtained, and for obtaining a target nucleic acid by cutting the section containing it out from the gel. Fujifilm Wako provides a variety of gel and related products for nucleic acid electrophoresis.
More Information
Nucleic acid electrophoresis is a method in which negatively charged nucleic acids are migrated through a carrier such as agarose gel or polyacrylamide gel and are separated according to their molecular weight (chain length), and is used for analysis and purification of nucleic acid fragments.
Principles of Nucleic Acid Electrophoresis
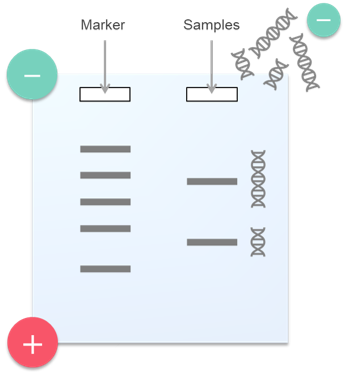
Nucleic acids are negatively charged due to their phosphate groups, and therefore can be induced to migrate through an electric field from a negative electrode toward a positive electrode. When this is carried out in a carrier such as agarose gel or polyacrylamide gel, the mobility of the nucleic acid fragments varies depending on the molecular weight: nucleic acid fragments with short chain lengths move easily through the gel, while those with long chain lengths move with more difficulty. As a result, nucleic acids can be separated according to their molecular weight (Fig. 1).
As double-stranded DNA (except for circular plasmid) generally has a linear structure, differences in molecular weight are simply reflected in mobility, whereas single-stranded DNA and RNA may have higher-order structures according to their nucleotide sequences, which affects their mobility. In order to separate single-stranded DNA or RNA molecules according to their molecular weight, a denaturing agent, such as urea or formamide, must be added to the gel or loading buffer to prevent the formation of higher-order structures. Conversely, experiments can be performed in which DNA or RNA is intentionally made to form a higher-order structure and is separated according to the difference in the structure.
How to Select Agarose or Polyacrylamide?
Agarose and polyacrylamide mainly used in nucleic acid electrophoresis. A summary of their features is provided below
Agarose
Agarose is a component of agar. When dissolved at high temperature and cooled, it becomes a gel with a network structure. Agarose used for nucleic acid electrophoresis is purified by removing other components without gelation ability, such as agaropectin.
Agarose gel has a larger network structure than polyacrylamide gel and is therefore used to separate relatively long nucleic acid fragments (50 to 20 kbp). Agarose can slide down an angled slab easily, and is not suitable for vertical slab systems such as those in which polyacrylamide gel is used. Agarose gel is commonly run in a submarine type electrophoresis unit in which the gel is set horizontally and is submerged in buffer.
Polyacrylamide
Polyacrylamide is a polymer of acrylamide, and bisacrylamide is used as a cross-linker. The gel network structure of polyacrylamide is finer than that of agarose and is suitable for the separation of short nucleic acid fragments (5 to 500 bp). When migration conditions are optimized, it is possible to detect a difference of a single base, based on the difference in migration rate.
Reagents Required for Electrophoresis
Running Buffer
Tris-acetate-EDTA (TAE) buffer or Tris-borate-EDTA (TBE) buffer is often used as running buffer for nucleic acid electrophoresis. TAE buffer has a high ability to separate long DNA fragments and TBE buffer has a high ability to separate short DNA fragments; therefore, TAE buffer is selected for agarose electrophoresis and TBE buffer is selected for polyacrylamide electrophoresis.
Loading buffer
Loading buffer is mixed with a sample when the sample is introduced (loaded) into a well in a gel. The roles of the loading buffer include (1) to increase the specific gravity of the sample to make it easier to load into the well, (2) to color the sample for visual confirmation of the sample loaded into the well, and (3) to monitor the progress of electrophoresis using the buffer as a dye.
Glycerol or Ficoll® PM400 is commonly used as a specific gravity additive for loading buffer. It is said that when Ficoll® is used, a narrow band can be obtained and “smiling” of bands is unlikely to occur compared with glycerol.
Bromophenol Blue (BPB), Xylene cyanol FF (XC), or Orange G is commonly used as a dye. When BPB is used, the dye produces a shadow that may be observed in a photograph. Although BPB does not have a large effect on experiments, it is necessary to use a loading buffer without BPB if presence of a dye shadow is a concern.
* Ficoll is a registered trademark of Cytiva
Molecular weight marker
A molecular weight marker, also called as DNA or RNA ladder, is a solution containing nucleic acid fragments for which the molecular weight (fragment size) is known. A molecular weight marker and a sample are electrophoresed simultaneously in different lanes, and the electrophoretic mobilities are compared, enabling estimation of the length of the nucleic acid in the sample. A molecular weight marker can be prepared by fragmenting a plasmid with specific restriction enzymes or by amplifying fragments of a defined length by PCR.
Nucleic acid staining reagent
A dye for which the fluorescence intensity increases when it is bound to nucleic acid is used for detection of nucleic acids electrophoresed in either agarose or polyacrylamide gel. Additional information about nucleic acid staining reagents can be found here.
References
- “Electrophoresis Q&A” ed.by Oto, M., Yodosha, Japan, (2004). (Japanese)
- “Principles and protocols for nucleic acid experiments by purpose” ed. by Hirao, I., Kurumizaka, H., Yodosha, Japan, (2011). (Japanese)
For research use or further manufacturing use only. Not for use in diagnostic procedures.
Product content may differ from the actual image due to minor specification changes etc.
If the revision of product standards and packaging standards has been made, there is a case where the actual product specifications and images are different.
The prices are list prices in Japan.Please contact your local distributor for your retail price in your region.



